n2p2 - Polynomial Symmetry Functions¶
This module introduces a new set of atomic environment descriptors for high-dimensional neural network potentials (HDNNPs) in n2p2. Polynomial symmetry functions [1] are designed to mimic closely the behavior of traditional Behler-Parrinello symmetry functions [2] but with a significantly reduced computational cost.
Purpose of Module¶
The symmetry functions proposed in the original work of Behler and Parrinello
[2] contain expressions of the form 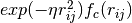 in
the innermost loop over all neighbors of atoms. Often the cutoff function
in
the innermost loop over all neighbors of atoms. Often the cutoff function
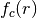 is chosen to be a cosine or hyperbolic tangent. Considering the
computational cost of these transcendental functions an alternative formulation
of symmetry functions based on polynomials like
is chosen to be a cosine or hyperbolic tangent. Considering the
computational cost of these transcendental functions an alternative formulation
of symmetry functions based on polynomials like
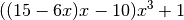
has been published recently [1]. Here, cheap polynomials are combined to form compact functions in the radial and angular domain which mimic the behavior of Behler-Parrinello type symmetry functions at a significantly reduced execution time. The benefits, benchmarks and many example applications are presented in great detail in [1].
This module’s changes to the n2p2 code comprise of new classes for different types of polynomial symmetry functions (PSFs), some helper classes and a redesign of the symmetry function caching mechanism:
Helper classes
CoreFunctionandCompactFunctionallow unified access to the compact function building blocks of PSFs.Six different types of PSFs were implemented in these classes:
SymFncCompRadSymFncCompAngnSymFncCompAngwSymFncCompRadWeightedSymFncCompAngnWeightedSymFncCompAngwWeighted
Here,
RadandAngn/Angwindicate radial and angular symmetry functions variants, respectively. The suffixWeightedrefers to an element weighting proposed in [3]. For each new class in this list also a symmetry function group [4] version was implemented, following the same naming scheme prefixed withSymGrp. See also this section of the n2p2 documentation for a more detailed description of the PSFs and their parameters.The computation of a set of descriptors allows the reuse of intermediate results across multiple symmetry functions with varying parameters. The previously existing cutoff function caching [4] of n2p2 was significantly improved. By overriding the
getCacheIdentifiers()member function eachSymFnc..class can provide identifier strings for required cache fields. TheMode::setupSymmetryFunctionCache()function collects the requirements of all symmetry functions and assigns cache positions in theAtom::Neighbor::cachearray.
Background Information¶
This module is based on n2p2, a C++ code for generation and application of neural network potentials used in molecular dynamics simulations. The source code and documentation are located here:
- n2p2 documentation: http://compphysvienna.github.io/n2p2/
- n2p2 source code: http://github.com/CompPhysVienna/n2p2
Building and Testing¶
The code changes from this module are already merged with the main repository of n2p2 (see pull request).
Because the introduction of a new set of symmetry function enhances the core
library of n2p2 several applications shipped with n2p2 will be affected by
the changes. The easiest way to test the new functionality is to run the examples
provided in these examples/nnp-predict/ folders which make use of PSFs:
Anisole_SCANDMABN_SCANEthylbenzene_SCAN
First, since the changes from this module are already merged with the main
repository of n2p2 (see pull request) it is sufficient to
download the latest version. Then,
compile the nnp-predict tool by running
make nnp-predict -j
in the src directory. Next, switch to one of the above example directories
and run the prediction tool:
../../../bin/nnp-predict 0
In the SETUP: SYMMETRY FUNCTIONS section of the output there should be
symmetry functions with type (column tp) between 20 and 25 which identifies
different variants
of PSFs. In addition, the section SETUP: SYMMETRY FUNCTION CACHE contains an
overview of the cache usage.
Regression testing is implemented in n2p2 and automatically performed upon submission of a pull request via Travis CI. The log file showing the successful pass of all tests for the specific pull request can be found here. The tests include the above prediction examples and also perform a comparison of analytic and numeric derivatives of symmetry functions.
Source Code¶
The easiest way to view the source code changes covered by this module is to use the GitHub pull request page. There, use the Files changed tab to review all changes.
| [2] | (1, 2) Behler, J. Atom-Centered Symmetry Functions for Constructing High-Dimensional Neural Network Potentials. J. Chem. Phys. 2011, 134 (7), 074106. |
| [4] | (1, 2) Singraber, A.; Behler, J.; Dellago, C. Library-Based LAMMPS Implementation of High-Dimensional Neural Network Potentials. J. Chem. Theory Comput. 2019, 15 (3), 1827–1840. |