E-CAM Verlet_list_for_ODE module¶
The Verlet_list_for_ODE module introduces
Verlet-list
for the rigid-body dynamics
Open-Dynamics-Engine software.
Purpose of Module¶
Rigid-body dynamics is useful for mechanical articulated systems. In addition, the tool allows the user to simulate complex shape and resolve excluded volume constraints. It is used in the industry of video games to accurately reproduce physics. However, a software like Open-Dynamics-Engine computes pairwise overlap every time-step. The engine starts with a partition of the space and then loops over all the blocks of partition. For each blocks it runs nested loops to check the overlaps between the objects inside the block.
The module implements external functions that can be used to compute the Verlet-list. Therefore, the user does not call the pairwise overlap check every time-step. He only needs to loop over the Verlet-list with the pairwise objects within a given cut-off distance. However, the Verlet-list has to be updated according to the displacement length of the objects.
The module can be used to speed-up the Open-Dynamics-Engine simulation of polymers and complex objects system.
We test the module with two examples: chromatin fiber and bacterial circular DNA.
Chromatin fiber is an assembly of DNA wrapped around
nucleosomes that compact the genome.
We model the DNA at the scale of 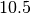 base-pair (one helix turn)
as an articulated system.
The nucleosomes is built with complex shape.
We run a Langevin dynamics
and check that our Verlet-list
implementation gives the same results
Open-Dynamics-Engine would give.
base-pair (one helix turn)
as an articulated system.
The nucleosomes is built with complex shape.
We run a Langevin dynamics
and check that our Verlet-list
implementation gives the same results
Open-Dynamics-Engine would give.
Background Information¶
You can find a detailed description on the Verlet_list_for_ODE GitLab repository.
Building and Testing¶
First of all you need to download and build the 0.16 version of Open-Dynamics-Engine. You can find the steps on the Verlet_list_for_ODE GitLab repository.
In order to compile the two examples (tests) I provide a template Makefile
you can find at the same location that the source code.
You need C++11 in order to use pseudo-random number generator.
It has OpenMP acceleration,
edit the Makefile to enable it. The example uses threads
from std as well.
Before the compilation you can clean the
previous build with make mrproper command.
The Verlet-list
implementation returns
the number of collisions that can be confronted
with the result returns by Open-Dynamics-Engine.
Source Code¶
The source code and more information can be find at Verlet_list_for_ODE GitLab repository.