Coarse-Graining: A Component of the Hierarchical Equilibration Strategy for Polymer Melts¶
The module is an implementation of the existing hierarchical strategy [1] for the equilibration of simple one-component polymer melts in ESPResSO++.
Purpose of Module¶
To study the properties of polymer melts by numerical simulations, equilibrated configurations must be prepared. However, the relaxation time for high molecular weight polymer melts is huge and increases, according to reptation theory, with the third power of the molecular weight. Hence, an effective method for decreasing the equilibration time is required. The hierarchical strategy pioneered in Ref. [1] is a particularly suitable way to do this. The present module provides a part of that method described below.
To decrease the relaxation time, microscopic monomers are coarse-grained (CG)
by mapping each subchain with 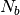 monomers onto a soft blob.
The CG system is then characterized by a much lower molecular weight and
thus is equilibrated quickly.
The present module provides a python script which performs this
coarse-graining procedure. The implementation detail is in following below.
monomers onto a soft blob.
The CG system is then characterized by a much lower molecular weight and
thus is equilibrated quickly.
The present module provides a python script which performs this
coarse-graining procedure. The implementation detail is in following below.
The microscopic configuration of
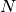 polymers consisted of
polymers consisted of 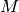 monomers is prepared. The system size
monomers is prepared. The system size 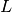 is determined by the number of density
is determined by the number of density 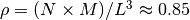 .
. 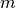 and
and 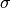 stands for the mass and the diameter of monomers.
stands for the mass and the diameter of monomers.The configuration of
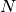 CG chain at
CG chain at 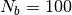 is generated from the microscopic configuration.
is generated from the microscopic configuration.The position of
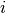 -th softblobs
-th softblobs 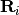 is determined by
is determined by 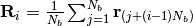 , where
, where 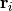 stands for the position of
stands for the position of 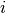 -th monomers.
-th monomers.The radius of gyration
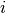 th softblobs
th softblobs 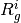 is also determined by
is also determined by 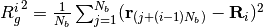 .
.The CG configuration is equilibrated by NVT MD simulation with mass of softblobs
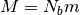 during the equilibration time
during the equilibration time 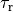 defined as
defined as 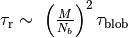 , where
, where 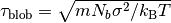 .
.Hence, CPU time
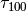 for softblobs with
for softblobs with 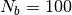 is estimated as
is estimated as 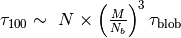 .
.After equilibrating a configuration, we continue to carry out MD simulation for adopting the snapshot which show ideal mean square internal distance (MSID)
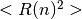 represented as
represented as 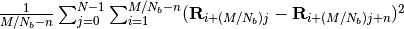 .
.Ideal MSID means the MSID of CG chains generated from fully equilibrated microscopic configurations.
A snapshot is captured in each
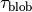 .
.
Background Information¶
The implementation of this module is based on ESPResSO++. You can learn about ESPResSO++ from the following links:
- ESPResSO++ documentation: http://espressopp.github.io/ESPResSo++.pdf
- ESPResSO++ source code: https://github.com/espressopp/espressopp
Building and Testing¶
Explanation of installation:
After installing this module, it can be tested according to the README file found under the following link:
Source Code¶
This module has been merged into ESPResSo++: