Dask-traj¶
For analysis of MD simulations MDTraj is a fast and commonly used analysis. However MDTraj has limitations, such as the requirement that the whole trajectory and result of the computation fits into memory. This module rewrites part of MDTraj to work with Dask in order to achieve out-of-memory computations, and combined with dask-distributed results in possible out-of-machine parallelization, essential for HPCs and a (surprising) speed-up even on a single machine.
Purpose of Module¶
Using MDTraj is a fast and easy way to analyze MD trajectories. However, MDTraj has a couple limitations:
- The whole trajectory needs to fit into memory, or gathering results becomes inconvenient
- The result of the computation also needs to fit into memory
- All processes need access to all the memory, preventing out-of-machine parallelization, and HPC scaling
Dask-traj solves all 3 limitations by rewriting the MDTraj functions to work with dask.arrays. This is done for both the trajectory and the computation functions. As dask.arrays know how to spill to disk, this lifts the requirement to fit into memory on both.
Together with dask-distributed it also allows the computation to be executed in a distributed way, which allows scaling out of a single machine. In preliminary tests this approach even leads to a speedup on a single machine, which is surprising as MDTraj is already a parallel code.
The splitting of everything in Dask-traj is done in the time-axis of the MD trajectory and as a lot of analysis is embarrassingly parallel, this leads to nice non-communicating compute graphs as shown here.
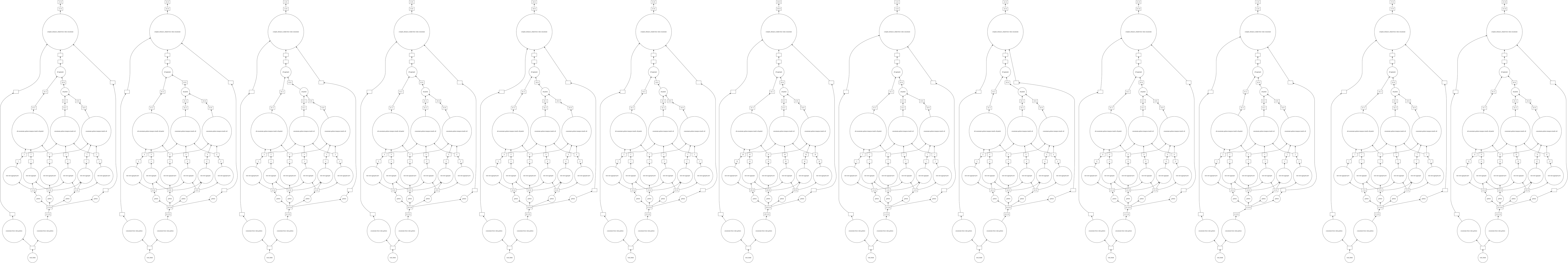
Current Limitations¶
One very important point of dask-traj is that we seek in the trajectory file.
So if your files are stored in a format that does not have an efficient seek
method, the loading of Trajectories will not get a speed-up, and might even be
slower than MDTraj.
Also, due to the way the code is written in MDTraj, only a subset of functions are available at the moment, but this will be expanded further in the future. If you have a use-case that requires the conversion of a MDTraj functionality, not yet present in dask-traj, please make an issue and I will focus on that.
Building and Testing¶
This code can be installed with conda using conda install -c dask_traj. To
install the specific version associated with this module, use conda install -c
conda-forge dask_traj==0.2.2
This code can also be installed with pip by running
pip install dask-traj
Finally, this code can also be installed by downloading the source code (see the Source
Code section below), and running python setup.py install from the root
directory.
Tests for this module can be run with pytest. Install pytest with pip
install pytest and then run the command py.test from within the
directory with the source code, or py.test --pyargs dask_traj from
anywhere after installation.
Examples¶
The examples require some extra dependencies to be installed, namely: * jupyter * distributed * python-graphviz
Which are all installable through conda and pip.
- An example on how to do analysis using Dask-traj can be found in dask-traj_example.ipynb
- An example on how to combine dask-traj with dask.distributed can be found in dask-traj_distributed example.ipynb
These examples can also be found in the examples directory in the source code. They can be run by
using jupyter notebook from that directory (see Jupyter notebook documentation at http://jupyter.org/ for more details)
Source Code¶
The source code for this module, and modules that build on it, is hosted at https://github.com/sroet/dask-traj. This module specifically includes everything up to and including release 0.2.2