E-CAM High Throughput Computing Library¶
E-CAM is interested in the challenge
of bridging timescales. To study molecular dynamics with atomistic detail, timesteps must be used on
the order of a femtosecond. Many problems in biological chemistry, materials science, and other
fields involve events that only spontaneously occur after a millisecond or longer (for example,
biomolecular conformational changes, or nucleation processes). That means that around 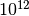 time
steps would be needed to see a single millisecond-scale event. This is the problem of “rare
events” in theoretical and computational chemistry.
time
steps would be needed to see a single millisecond-scale event. This is the problem of “rare
events” in theoretical and computational chemistry.
Modern supercomputers are beginning to make it possible to obtain trajectories long enough to observe some of these processes, but to fully characterize a transition with proper statistics, many examples are needed. In order to obtain many examples the same application must be run many thousands of times with varying inputs. To manage this kind of computation a task scheduling high throughput computing (HTC) library is needed. The main elements of mentioned scheduling library are: task definition, task scheduling and task execution.
While traditionally an HTC workload is looked down upon in the HPC space, the scientific use case for extreme-scale resources exists and algorithms that require a coordinated approach make efficient libraries that implement this approach increasingly important in the HPC space. The 5 Petaflop booster technology of JURECA is an interesting concept with respect to this approach since the offloading approach of heavy computation marries perfectly to the concept outlined here.
Purpose of Module¶
This module is the first in a sequence that will form the overall capabilities of the library. In particular this module deals with creating a set of decorators to wrap around the Dask-Jobqueue Python library, which aspires to make the development time cost of leveraging it lower for our use cases.
Background Information¶
The initial motivation for this library is driven by the ensemble-type calculations that are required in many scientific fields, and in particular in the materials science domain in which the E-CAM Centre of Excellence operates. The scope for parallelisation is best contextualised by the Dask documentation:
A common approach to parallel execution in user-space is task scheduling. In task scheduling we break our program into many medium-sized tasks or units of computation, often a function call on a non-trivial amount of data. We represent these tasks as nodes in a graph with edges between nodes if one task depends on data produced by another. We call upon a task scheduler to execute this graph in a way that respects these data dependencies and leverages parallelism where possible, multiple independent tasks can be run simultaneously.
Many solutions exist. This is a common approach in parallel execution frameworks. Often task scheduling logic hides within other larger frameworks (Luigi, Storm, Spark, IPython Parallel, and so on) and so is often reinvented.
Dask is a specification that encodes task schedules with minimal incidental complexity using terms common to all Python projects, namely dicts, tuples, and callables. Ideally this minimum solution is easy to adopt and understand by a broad community.
While we were attracted by this approach, Dask did not support task-level parallelisation (in particular multi-node tasks). We researched other options (including Celery, PyCOMPSs, IPyParallel and others) and organised a workshop that explored some of these (see https://www.cecam.org/workshop-0-1650.html for further details).
Building and Testing¶
The library is a Python module and can be installed with
python setup.py install
More details about how to install a Python package can be found at, for example, Install Python packages on the research computing systems at IU
To run the tests for the decorators within the library, you need the pytest Python package. You can run all the
relevant tests from the jobqueue_features directory with
pytest tests/test_decorators.py
Examples of usage can be found in the examples directory.
Source Code¶
The latest version of the library is available on the jobqueue_features GitHub repository, the file specific to this module is decorators.py.
(The code that was originally created for this module can be seen in the specific commit 4590a0e427112f which can be found in the original private repository of the code.)